
Overview of next generation sequencing
In 1972, Sanger sequencing was first developed with dideoxy chain termination for DNA sequencing approach. This method has been widely used to determine the DNA sequences with more than 99.9% accuracy. However, it is inefficient because it can only sequence one sample at a time and the cost per base is expensive. Therefore, new high-throughput sequencing technologies have been developed to reduce cost and increase sample acquired per run. These technologies are described as next-generation sequencing (NGS). In 2005, Roche 454 GS-20 sequencing, the first NGS platform, was released on the market, which can produce 20 million of sequences. The timeline where different NGS technologies have been developed during the last 15 years is shown in Figure 1 (Gupta AK, Gupta U, 2020).
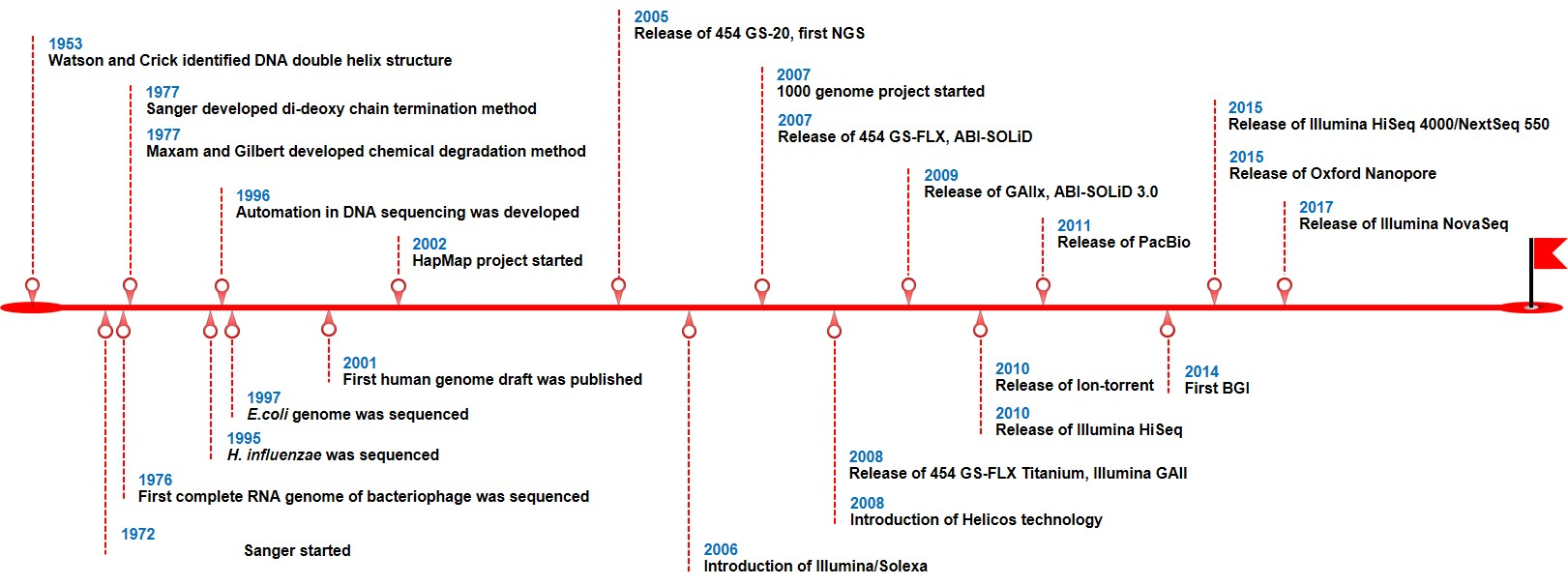
NGS technologies are capable of sequencing in an extensively parallel ways, which can generate massive amount of information (millions of sequences) very rapidly at once. NGS technologies included Roche 454, Illumina, and ABI-SOLiD platforms. NGS is currently considered as a gold standard sequencing for clinical and biological investigations, diagnostics, and evolutionary analysis in plant and animal researches.
References :
Gupta AK, Gupta U. Next generation sequencing and its applications. Anim Biotechnol 2020:395–421. doi:10.1016/b978-0-12-811710-1.00018-5